AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog
 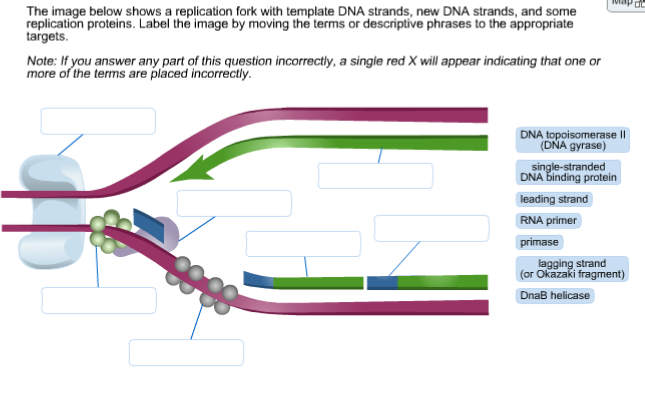
This continuously synthesized strand is known as the leading strand. One strand, which is complementary to the 3' to 5' parental DNA strand, is synthesized continuously towards the replication fork because the polymerase can add nucleotides in this direction. As we know, the DNA double helix is anti-parallel that is, one strand is in the 5' to 3' direction and the other is oriented in the 3' to 5' direction. The gap is then filled by a polymerase (/). Then, DNA polymerase begins to remove the RNA primers. There is a leading and a lagging strand for each of the two replication forks. DNA polymerase can only extend in the 5' to 3' direction, which poses a slight problem at the replication fork. Both leading and Okazaki fragments of lagging strands are synthesized from 5´ 3´ direction. The replication fork moves at the rate of 1000 nucleotides per second. This second DNA polymerase then synthesizes the remainder of each Okazaki fragment with the help of a clamp protein (Figure 5-28). (credit: modification of work by Mariana Ruiz Villareal) The polymerase begins each Okazaki fragment on the lagging strand with RNA and then extends the RNA primer with a short length of DNA, before passing the 3 end of this primer to a second enzyme, DNA polymerase. DNA ligase seals the gaps between the Okazaki fragments, joining the fragments into a single DNA molecule. DNA polymerase I replaces the RNA primer with DNA. Iterative cycles between Polymerase (Pol) and Flap endonuclease. c. b.) On the Lagging strand and are processed and joined by DNA ligase to form a continuous strand of DNA. On the leading strand, DNA is synthesized continuously, whereas on the lagging strand, DNA is synthesized in short stretches called Okazaki fragments. The final steps of lagging strand synthesis induce maturation of Okazaki fragments via removal of the RNA primers and ligation. During replication of DNA, Okazaki fragments are: a.) On the Lagging strand and are made up of RNA primer and short DNA sequences. DNA polymerase III uses this primer to synthesize the daughter DNA strand. DNA polymerase in eukaryotes and DNA polymerase I in prokaryotes then extends it close to the replication fork. Single-strand binding proteins bind to the single-stranded DNA to prevent the helix from re-forming. RNA primers made by primase are the starting point for synthesizing each Okazaki fragment. Topoisomerase breaks and reforms DNA’s phosphate backbone ahead of the replication fork, thereby relieving the pressure that results from this supercoiling. The DNA tends to become more highly coiled ahead of the replication fork. \): A replication fork is formed when helicase separates the DNA strands at the origin of replication. 
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
